Single-fidelity VeBRNN¤
This notebook guidance on how to train VeBRNN of the MF-VeBRNN repo.
# import packages
import torch
import matplotlib.pyplot as plt
from torch import nn
import warnings
warnings.filterwarnings("ignore")
from matplotlib.ticker import ScalarFormatter
from VeBNN.networks import MeanNet, GammaVarNet
from MFVeBRNN.method.vebnn_trainer import VeBRNNTrainer
from MFVeBRNN.dataset.load_dataset import SingleFidelityDataset
import numpy as np
Load the plasticity law discovery datset¤
The plasticity law discovery dataset can be loaded by the class SingleFidelityDataset.
dataset = SingleFidelityDataset(train_data_path = "lf_dns_sve_0d1.pickle",
id_ground_truth=True,
id_test_data_path="hf_dns_rve_0d1_gt.pickle",
id_ground_truth_data_path="lf_dns_sve_0d1_gt.pickle",
ood_ground_truth=True,
ood_test_data_path="hf_dns_rve_0d125_gt.pickle",
ood_ground_truth_data_path='lf_dns_sve_0d125_gt.pickle',)
dataset.get_train_val_split(num_train=100,num_val=100)
=============================================================
The dataset is loaded successfully.
Number of training samples: 2981
Number of in-distribution test samples: 99
Number of out-of-distribution test samples: 99
=============================================================
Define a GRU network¤
Since the history dependent constitutive law is has recurrent data structure, we need a recurrent neural network for this task.
class SimpleGRU(nn.Module):
def __init__(self,
input_size: int,
hidden_size: int,
num_layers: int,
output_size: int,
bias: bool = True):
super().__init__()
self.gru = nn.GRU(input_size=input_size,
hidden_size=hidden_size,
num_layers=num_layers,
bias=bias,
batch_first=True,
)
self.h2y = nn.Linear(hidden_size, output_size)
def forward(self, x, hx=None):
out, _ = self.gru(x, hx) # (B, T, hidden_size)
y = self.h2y(out) # (B, T, output_size)
return y
Training setup¤
# define the mean and variance networks
mean_network_ = SimpleGRU(input_size=3,
hidden_size=128,
num_layers=2,
output_size=3)
mean_network = MeanNet(net=mean_network_,
prior_mu=0.0,
prior_sigma=1.0)
# define the variance network
var_network_ = SimpleGRU(input_size=3,
hidden_size=4,
num_layers=1,
output_size=6)
var_network = GammaVarNet(net=var_network_,
prior_mu=0.0,
prior_sigma=1.0)
# set up the configuration for the VeBNN trainer
trainer = VeBRNNTrainer(mean_net=mean_network,
var_net=var_network,
device=torch.device("cuda" if torch.cuda.is_available() else "cpu"),
job_id=1)
# ====== configs (kept small so test is fast) ======
init_config = {
"loss_name": "MSE",
"optimizer_name": "Adam",
"lr": 1e-3,
"weight_decay": 1e-6,
"num_epochs": 1000, # warm-up epochs
"batch_size": 200,
"verbose": False,
"print_iter": 50,
"split_ratio": 0.8,
}
var_config = {
"optimizer_name": "Adam",
"lr": 1e-3,
"num_epochs": 1000,
"batch_size": 200,
"verbose": True,
"print_iter": 50,
"early_stopping": False,
"early_stopping_iter": 100,
"early_stopping_tol": 1e-4,
}
sampler_config = {
"sampler": "pSGLD", # must match your VeBNN.samplers names
"lr": 1e-3,
"gamma": 0.9999,
"num_epochs": 2000, # SGMCMC epochs
"mix_epochs": 10, # thinning interval
"burn_in_epochs": 500,
"batch_size": 200,
"verbose": False,
"print_iter": 100,
}
# ====== run cooperative training ======
# For a quick test, iteration=2 is enough to see if everything works.
trainer.cooperative_train(
x_train=dataset.x_train,
y_train=dataset.y_train,
iteration=1,
init_config=init_config,
var_config=var_config,
sampler_config=sampler_config,
delete_model_raw_data=True, # delete temporary folder after training
)
=========================================================
Step 2: Train for the variance network, iteration 1
Epoch/Total: 0/1000, Gamma NLL: -3.280e+04, neg log prior: 1.323e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 50/1000, Gamma NLL: -5.184e+04, neg log prior: 1.324e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 100/1000, Gamma NLL: -7.267e+04, neg log prior: 1.328e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 150/1000, Gamma NLL: -9.174e+04, neg log prior: 1.335e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 200/1000, Gamma NLL: -1.041e+05, neg log prior: 1.343e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 250/1000, Gamma NLL: -1.114e+05, neg log prior: 1.351e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 300/1000, Gamma NLL: -1.159e+05, neg log prior: 1.357e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 350/1000, Gamma NLL: -1.189e+05, neg log prior: 1.363e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 400/1000, Gamma NLL: -1.209e+05, neg log prior: 1.368e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 450/1000, Gamma NLL: -1.223e+05, neg log prior: 1.373e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 500/1000, Gamma NLL: -1.233e+05, neg log prior: 1.377e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 550/1000, Gamma NLL: -1.241e+05, neg log prior: 1.381e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 600/1000, Gamma NLL: -1.247e+05, neg log prior: 1.385e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 650/1000, Gamma NLL: -1.251e+05, neg log prior: 1.389e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 700/1000, Gamma NLL: -1.255e+05, neg log prior: 1.392e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 750/1000, Gamma NLL: -1.258e+05, neg log prior: 1.395e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 800/1000, Gamma NLL: -1.261e+05, neg log prior: 1.399e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 850/1000, Gamma NLL: -1.263e+05, neg log prior: 1.402e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 900/1000, Gamma NLL: -1.265e+05, neg log prior: 1.405e+02, log marginal likelihood: 0.000e+00
Epoch/Total: 950/1000, Gamma NLL: -1.268e+05, neg log prior: 1.409e+02, log marginal likelihood: 0.000e+00
Finished training the variance network
===========================================
Step 3: Train for the mean network with SGMCMC
Finished training the Bayesian mean network
============================================
Create model data folder to save the temporary models
=========================================================
Finished training the model
Delete the model data folder to free space
Get VeRNN's prediction¤
# get the prediction from the bnn
pred, var_epistemic = trainer.bayes_predict(x=dataset.x_id_gt_scaled)
# for aleatoric uncertainty
var_aleatoric= trainer.aleatoric_variance_predict(
x=dataset.x_id_gt_scaled)
# convert to cpu
pred = pred.cpu().detach()
var_epistemic = var_epistemic.cpu().detach()
var_aleatoric = var_aleatoric.cpu().detach()
# set the
# scale the prediction back
pred = pred * dataset.Y_std + dataset.Y_mean
var_epistemic = var_epistemic * dataset.Y_std ** 2
var_aleatoric = var_aleatoric * dataset.Y_std ** 2
# save the model for later use
torch.save(trainer, "vebrnn_model.pth")
formatter = ScalarFormatter(useMathText=True)
formatter.set_scientific(True)
formatter.set_powerlimits((-1, 1))
def brnn_prediction_plot(
x_test: torch.Tensor,
y_test: torch.Tensor,
y_pred_mean: torch.Tensor,
y_pred_var: torch.Tensor,
y_pred_aleatoric: torch.Tensor,
index: int,
fig_name="rnn_prediction",
save_fig: bool = False,
y_test_var: torch.Tensor = None,
) -> None:
strain = x_test.cpu().detach().numpy()
stress = y_test.cpu().detach().numpy()
y_pred_mean = y_pred_mean.cpu().detach().numpy()
if y_pred_var is not None:
y_pred_var = y_pred_var.cpu().detach().numpy()
if y_test_var is not None:
y_test_var = y_test_var.cpu().detach().numpy()
if y_pred_aleatoric is not None:
y_pred_aleatoric = y_pred_aleatoric.cpu().detach().numpy()
fig, ax = plt.subplots(2, 3, figsize=(12, 5))
ax[0, 0].plot(strain[index, :, 0], "-", color="#0077BB", linewidth=2)
ax[0, 0].set_ylabel(ylabel=r"$E_{11}$", fontsize=12)
ax[0, 0].yaxis.set_major_formatter(formatter)
ax[0, 1].plot(strain[index, :, 1], "-", color="#0077BB", linewidth=2)
ax[0, 1].set_ylabel(ylabel=r"$E_{12}$", fontsize=12)
ax[0, 1].yaxis.set_major_formatter(formatter)
ax[0, 2].plot(strain[index, :, 2], "-", color="#0077BB", linewidth=2)
ax[0, 2].set_ylabel(ylabel=r"$E_{22}$", fontsize=12)
ax[0, 2].yaxis.set_major_formatter(formatter)
# plot the stress
ax[1, 0].plot(stress[index, :, 0], "-", linewidth=2, color="#0077BB")
if y_test_var is not None:
ax[1, 0].fill_between(
range(len(stress[index, :, 0])),
y1=y_test[index, :, 0] - 2 * np.sqrt(y_test_var[index, :, 0]),
y2=y_test[index, :, 0] + 2 * np.sqrt(y_test_var[index, :, 0]),
edgecolor="none",
color="#0077BB",
alpha=0.3,
)
ax[1, 0].plot(y_pred_mean[index, :, 0], "-", color="#CC3311", linewidth=2)
if y_pred_var is not None:
ax[1, 0].fill_between(
range(len(stress[index, :, 0])),
y1=y_pred_mean[index, :, 0] - 2 * np.sqrt(y_pred_var[index, :, 0]),
y2=y_pred_mean[index, :, 0] + 2 * np.sqrt(y_pred_var[index, :, 0]),
edgecolor="none",
color="#EE7733",
alpha=0.5,
)
# plot the aleatoric uncertainty
if y_pred_aleatoric is not None:
ax[1, 0].fill_between(
range(len(stress[index, :, 0])),
y1=y_pred_mean[index, :, 0] - 2 *
np.sqrt(y_pred_aleatoric[index, :, 0]),
y2=y_pred_mean[index, :, 0] + 2 *
np.sqrt(y_pred_aleatoric[index, :, 0]),
alpha=0.6,
color="gray",
edgecolor="none",
)
ax[1, 0].set_xlabel(xlabel="Time step", fontsize=12)
ax[1, 0].set_ylabel(ylabel=r"$\sigma_{11}$ (MPa)", fontsize=12)
ax[1, 0].yaxis.set_major_formatter(formatter)
ax[1, 1].plot(
stress[index, :, 1],
"-",
linewidth=2,
color="#0077BB",
)
if y_test_var is not None:
ax[1, 1].fill_between(
range(len(stress[index, :, 1])),
y1=y_test[index, :, 1] - 2 * np.sqrt(y_test_var[index, :, 1]),
y2=y_test[index, :, 1] + 2 * np.sqrt(y_test_var[index, :, 1]),
edgecolor="none",
color="#0077BB",
alpha=0.3,
)
ax[1, 1].plot(
y_pred_mean[index, :, 1],
"-",
color="#CC3311",
linewidth=2,
)
if y_pred_var is not None:
ax[1, 1].fill_between(
range(len(stress[index, :, 1])),
y1=y_pred_mean[index, :, 1] - 2 * np.sqrt(y_pred_var[index, :, 1]),
y2=y_pred_mean[index, :, 1] + 2 * np.sqrt(y_pred_var[index, :, 1]),
color="#EE7733",
edgecolor="none",
alpha=0.5,
)
# plot the aleatoric uncertainty
if y_pred_aleatoric is not None:
ax[1, 1].fill_between(
range(len(stress[index, :, 1])),
y1=y_pred_mean[index, :, 1] - 2 *
np.sqrt(y_pred_aleatoric[index, :, 1]),
y2=y_pred_mean[index, :, 1] + 2 *
np.sqrt(y_pred_aleatoric[index, :, 1]),
alpha=0.3,
color="gray",
edgecolor="none",
)
ax[1, 1].set_ylabel(ylabel=r"$\sigma_{12}$ (MPa)", fontsize=12)
ax[1, 1].set_xlabel(xlabel="Time step", fontsize=12)
ax[1, 1].yaxis.set_major_formatter(formatter)
if y_test_var is not None:
ax[1, 2].plot(
stress[index, :, 2], "-", linewidth=2, color="#0077BB", label="Ground Truth Mean"
)
ax[1, 2].fill_between(
range(len(stress[index, :, 2])),
y1=y_test[index, :, 2] - 2 * np.sqrt(y_test_var[index, :, 2]),
y2=y_test[index, :, 2] + 2 * np.sqrt(y_test_var[index, :, 2]),
edgecolor="none",
color="#0077BB",
alpha=0.5,
label="Ground Truth 95% CI",
)
else:
ax[1, 2].plot(
stress[index, :, 2], "-", linewidth=2, color="#0077BB", label="Test Data"
)
ax[1, 2].plot(
y_pred_mean[index, :, 2], "-", color="#CC3311", linewidth=2, label="Pred. Mean"
)
if y_pred_var is not None:
ax[1, 2].fill_between(
range(len(stress[index, :, 2])),
y1=y_pred_mean[index, :, 2] - 2 * np.sqrt(y_pred_var[index, :, 2]),
y2=y_pred_mean[index, :, 2] + 2 * np.sqrt(y_pred_var[index, :, 2]),
alpha=0.3,
color="#EE7733",
edgecolor="none",
label="Pred. 95% CI Total",
)
# plot the aleatoric uncertainty
if y_pred_aleatoric is not None:
ax[1, 2].fill_between(
range(len(stress[index, :, 2])),
y1=y_pred_mean[index, :, 2] - 2 *
np.sqrt(y_pred_aleatoric[index, :, 2]),
y2=y_pred_mean[index, :, 2] + 2 *
np.sqrt(y_pred_aleatoric[index, :, 2]),
alpha=0.3,
color="gray",
edgecolor="none",
label="Pred. 95% CI Aleatoric",
)
ax[1, 2].set_xlabel(xlabel="Time step", fontsize=12)
ax[1, 2].set_ylabel(ylabel=r"$\sigma_{22}$ (MPa)", fontsize=12)
ax[1, 2].legend(fontsize=8, loc="best", frameon=False)
ax[1, 2].yaxis.set_major_formatter(formatter)
# set the line width and font size for the axises
for i in range(2):
for j in range(3):
ax[i, j].tick_params(axis="both", which="major", labelsize=12)
# set linewidth
for axis in ["top", "bottom", "left", "right"]:
ax[i, j].spines[axis].set_linewidth(1.5)
# set the spaces for subplots
plt.subplots_adjust(wspace=0.32, hspace=0.25)
if save_fig:
# plt.savefig(f"{fig_name}.pdf", dpi=300, bbox_inches="tight")
plt.savefig(f"{fig_name}.png", dpi=300, bbox_inches="tight")
else:
plt.show()
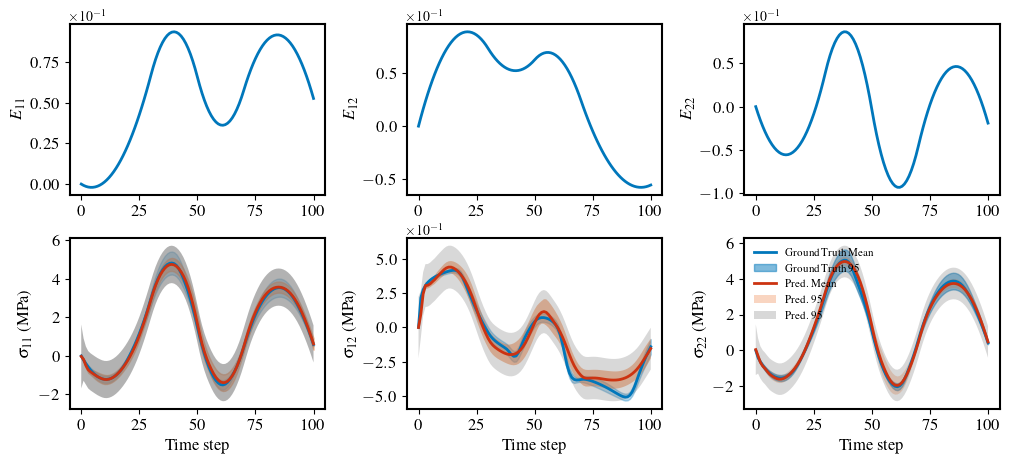
brnn_prediction_plot(
x_test=dataset.x_id_gt,
y_test=dataset.y_id_gt_mean,
y_pred_mean=pred,
y_pred_var=var_epistemic,
y_pred_aleatoric=var_aleatoric,
index=1,
save_fig=False,
y_test_var=dataset.y_id_gt_var,
)